Simulate. Optimize. Process.
Complete protein processing simulation — from raw seed to fractionated flour. Pretreatment, milling, and air classification in one powerful desktop application.
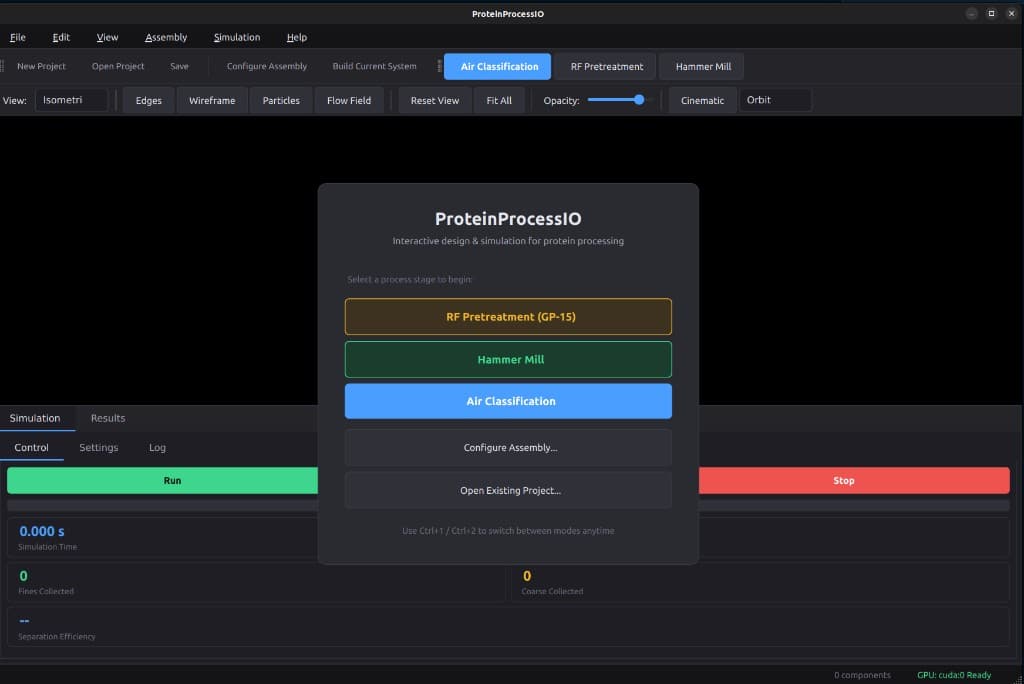
Complete Processing Pipeline
Simulate the entire protein fractionation process. Transfer data seamlessly between stages with automatic parameter mapping.
Powerful Features
Coupled multi-physics solvers, GPU-accelerated kernels, and experimentally validated models — from whole seed conditioning to micron-scale flour separation.
Coupled Multi-Physics Engine
9-Step LoopNine-step simulation loop coupling RF electromagnetic fields, heat conduction, moisture diffusion, and dielectric property updates. FDM Laplace solver with 100+ Lagrangian tracers for seed-level temperature and moisture tracking.
GPU-Accelerated Computation
CUDA/WarpEight NVIDIA Warp GPU kernels with persistent memory allocation and batched launches — zero per-step allocations. Automatic CUDA/CPU device detection with adaptive CFL-based timestep control.
Interactive 3D Digital Twin
40+ PartsOver 40 parametric components with three cinematic camera modes — orbit, guided showcase tours, and spiral flythrough. Live particle rendering, mechanical animations, and per-component opacity controls.
Three-Stage Pipeline
End-to-EndRF pretreatment, hammer milling, and air classification linked with automatic outlet-to-inlet state mapping. Multi-pass recirculation with attrition modeling and full mass balance tracking across stages.
Multi-Objective Optimization
5D ScoringDerringer-Suich desirability scoring across five dimensions — thermal treatment, LOX inactivation, protein preservation, moisture retention, and energy efficiency. Grid search and gradient-based optimization via Warp Tape.
Advanced Analytics & Export
VTK/CSVInteractive PSD charts with hover tooltips and log-scale toggle, real-time KPI dashboards with radial gauges, and dual-scale time-series plots. Export to VTK for 3D field post-processing, plus CSV, JSON, and NumPy formats.
Partners & funding
Developed at the Sustainable Agrifood Systems Engineering Lab (SASEL) at McGill University. Principal Investigator: Dr. Ebenezer Miezah Kwofie. Funded by the National Research Council of Canada (NRC).
Ready to optimize your process?
Download ProteinProcessIO today and start simulating your protein processing operations. Free for research and academic use.


